

From the fundamental perspectives of biochemistry and molecular engineering, this article systematically analyzes the complete workflow of recombinant protein production, from gene sequence design to the acquisition of high-purity samples. The discussion focuses on the impact of codon optimization on translational elongation and co-translational folding, examines the molecular basis of protein stability and protease inhibition during cell lysis, and explains the physicochemical principles underlying affinity chromatography, ion exchange chromatography, and size exclusion chromatography. In addition, criteria for evaluating protein monodispersity using SDS-PAGE and light-scattering techniques are discussed, providing a theoretical framework for preparing structural-biology-grade protein samples.
In modern biotechnology and structural biology, recombinant protein expression and purification represent the critical interface between genetic information and experimentally tractable molecular entities. For studies aimed at X-ray crystallography or cryo-electron microscopy, the mere presence of a protein band is insufficient. The true objective is to obtain a sample that is conformationally homogeneous, chemically pure, and folded in a native-like state. From this perspective, protein production is not a routine workflow, but a systems-level process that requires precise control of molecular interactions and energetic landscapes at every stage.
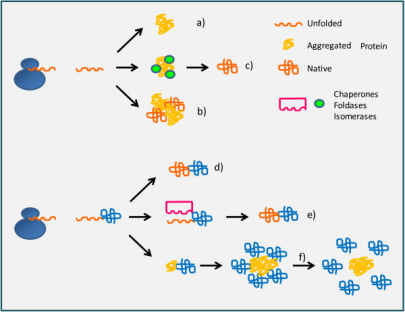
Recombinant protein production begins with gene sequence design. Although the genetic code is degenerate, synonymous codons are not used with equal frequency across organisms. This codon usage bias directly influences translational elongation kinetics. When a heterologous gene contains a high proportion of rare codons for the chosen host, ribosomes are prone to stalling due to limited availability of the corresponding tRNAs. Such pauses can expose hydrophobic segments of nascent polypeptides before proper folding occurs, increasing the likelihood of misfolding or proteolytic degradation.
Codon optimization aligns the coding sequence with the host’s tRNA abundance profile, promoting a smoother elongation rate and facilitating co-translational folding. In parallel, transcriptional strength and induction conditions must be tuned to the folding capacity of the cellular environment. Excessively rapid synthesis can overwhelm molecular chaperone systems, favoring aggregation over productive folding. At the molecular level, expression efficiency and solubility therefore exist in a dynamic balance that must be respected for successful protein production.
Following expression, the target protein remains embedded in a highly complex cellular matrix and must be released through cell lysis. Whether achieved by mechanical disruption, enzymatic digestion, or pressure-based methods, lysis instantaneously abolishes intracellular compartmentalization and liberates large quantities of endogenous proteases. These enzymes readily attack flexible or exposed regions of recombinant proteins and represent a major threat to sample integrity.
To mitigate proteolysis, lysis is typically performed at low temperature in the presence of broad-spectrum protease inhibitors. In addition, chromosomal DNA released during lysis dramatically increases solution viscosity, impairing downstream clarification and chromatography. Controlled degradation of nucleic acids using nucleases, followed by high-speed centrifugation, is therefore essential to obtain a clarified lysate suitable for subsequent purification steps.
The first purification stage is usually a capture step designed to rapidly enrich the target protein from a complex mixture. Affinity chromatography achieves this goal through specific molecular recognition. In the case of polyhistidine-tagged proteins, imidazole side chains coordinate reversibly with immobilized metal ions such as Ni²⁺ or Co²⁺, enabling selective binding under defined buffer conditions.
After non-specifically bound contaminants are removed, competitive elution disrupts the coordination interaction, yielding concentrated protein with substantial purity gains. Other affinity systems operate on analogous principles of selective binding, differing mainly in the nature and strength of the recognition interaction. The unifying feature of affinity chromatography is its ability to dramatically reduce sample complexity in a single step.
Following capture, ion exchange chromatography (IEX) is commonly employed to further refine sample purity. This technique exploits differences in protein surface charge, which are governed by amino acid composition, three-dimensional structure, and post-translational modifications. Each protein has a characteristic isoelectric point (pI) at which its net charge is zero.
When the buffer pH deviates from the pI, proteins acquire a net positive or negative charge and can bind to oppositely charged resins. Gradual increases in ionic strength weaken electrostatic interactions, allowing proteins to elute according to their charge properties. Because surface charge is sensitive to subtle structural variations, IEX can resolve closely related species, including modified or partially misfolded forms of the same protein.
Size exclusion chromatography (SEC) typically represents the final purification step and serves both as a polishing technique and a quality control assay. SEC separates molecules based on their hydrodynamic volume rather than chemical affinity. Porous matrix beads allow smaller species to explore a larger internal volume, delaying their elution relative to larger molecules.
For a well-behaved recombinant protein, SEC should yield a single, symmetric peak corresponding to a monodisperse population. Asymmetric peaks, shoulders, or early-eluting species often indicate aggregation or oligomerization. SDS-PAGE analysis of collected fractions provides information on molecular integrity, while light-scattering techniques further quantify size homogeneity in solution. For structural biology applications, monodispersity is a prerequisite for crystallization and high-resolution cryo-EM data collection.
Recombinant protein production is a multidisciplinary process that integrates molecular biology, biochemistry, and physical separation science. From translational kinetics dictated by gene design to the physicochemical principles governing chromatographic separation, each stage exerts a direct influence on final sample quality. Only by understanding and respecting these underlying mechanisms can researchers consistently obtain protein preparations suitable for high-resolution structural analysis.