

This article summarizes key determinants underlying high-performance CHO stable cell line formation from the perspectives of cell biology and molecular engineering. It focuses on (i) integration architectures of exogenous expression units in host chromosomes and associated position effects, (ii) copy-number dynamics and expression burden shaped by DHFR- and GS-based selection systems, and (iii) the impact of epigenetic states on long-term expression persistence. In addition, it outlines how standardized sample handling and analytical metrics support evaluation of product molecular consistency and heterogeneity distributions.
CHO cells are widely used mammalian hosts for complex recombinant protein expression due to their compatibility with suspension culture, robust secretory capacity, and glycosylation processing. In this context, “high performance” generally refers to sustained high secretion rates with reproducible expression behavior, whereas “stability” emphasizes persistence of transcriptional activity and product characteristics across serial passaging and across culture batches. These attributes are not independent; they are co-determined by genome configuration, cellular state, and processing burden imposed by exogenous expression.
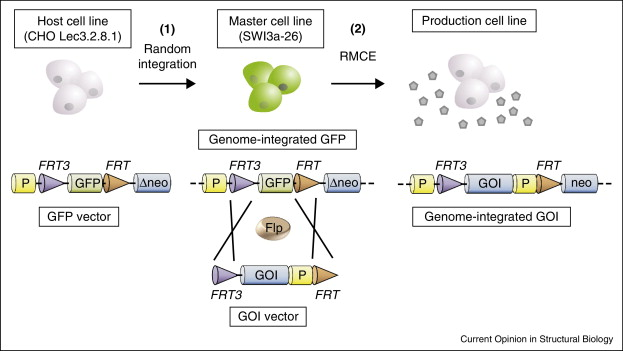
A defining event in stable cell line development is the chromosomal placement and maintenance of the exogenous expression unit. After delivery of an expression vector or plasmid DNA, exogenous sequences can adopt distinct genomic configurations, including differences in insertion site, local copy arrangement, and structural organization of the integrated cassette. Random integration frequently introduces position effects: transcriptional accessibility varies with chromatin context and neighboring regulatory features, leading to clone-to-clone differences in expression magnitude and differences in temporal drift during passaging. These effects reflect variation in local chromatin openness, proximity to enhancers or repressive elements, and susceptibility to progressive silencing.
By contrast, approaches that favor targeted integration or transposon-mediated integration are commonly discussed in terms of their ability to increase the probability of placement within transcriptionally permissive regions and to support more durable inheritance of the integrated unit. The mechanistic focus is therefore the sustained accessibility of regulatory elements and the stability of the integrated architecture, rather than maximizing short-term expression alone. In research analyses, integration-related variation is typically interpreted using both genetic-level measurements and expression-level measurements. For example, qPCR can provide quantitative support for copy-structure comparisons, and these data are considered alongside transcript- or protein-level changes to assess how integration architecture contributes to persistence of expression.
Within CHO stable cell line workflows, GS- and DHFR-related selection strategies are commonly used to couple cell survival to the maintenance of an exogenous expression unit, thereby enriching for cells that retain functional expression cassettes and, under certain conditions, exhibit copy number amplification. Importantly, increased copy number does not necessarily translate linearly into increased product output. The resulting phenotype depends on the capacity of the cell to sustain transcription, translation, folding, assembly, and secretion under the increased burden created by higher mRNA supply and higher protein synthesis rates.
Copy number amplification can increase transcript abundance, but it simultaneously elevates endoplasmic reticulum folding and assembly load, potentially affecting cellular physiology and shaping product heterogeneity distributions. Consequently, high performance is better described as a state in which exogenous expression flux is compatible with the host’s processing capacity over time, rather than as a single parameter such as copy number or a short-term secretion peak. For clone comparison, expression quantity metrics are therefore interpreted together with quality-attribute-related metrics to avoid over-reliance on a single productivity indicator when assessing suitability of a clone for further study.
A common explanation for expression decline during serial passaging in CHO cells involves epigenetic silencing. Following genomic integration, local DNA methylation patterns, histone modifications, and chromatin structural changes can reduce promoter accessibility and gradually suppress transcription. This phenomenon can occur even when the integrated sequence remains physically present, reflecting regulatory rather than structural loss.
To increase expression persistence, expression constructs may incorporate cis-regulatory elements intended to maintain a more open chromatin environment around the integration locus, such as MARs or UCOEs. The technical rationale is to reduce the probability of progressive silencing and to promote more stable transcriptional initiation over many generations. In mechanistic discussions, stability is best defined by consistency of expression behavior after extended passaging, supported by concordant evidence across genetic-level metrics, transcription-associated metrics, and protein-associated metrics. Short-term measurements alone are not sufficient to infer long-term performance because epigenetic drift is time-dependent and may manifest after multiple generations.
After establishment of stable cell lines, clonal variation is commonly observed across expression level, growth behavior, and product molecular population distributions. Although single-cell derivation and clone confirmation may involve tools such as flow cytometry, the central technical question is how to construct a consistent evaluation scheme so that different clones and different batches are compared under comparable conditions.
Assessment of product molecular consistency requires standardized sample handling and standardized analytical workflows. This ensures that observed differences in integrity, conformational state, or aggregation-associated features are attributable to biological variation rather than inconsistent sample preparation. Common analytical items include SDS-PAGE for evaluating integrity and principal molecular-weight features, and SEC for characterizing aggregation-related distributions and solution-state behavior. When needed, genetic quantification such as qPCR can support localization of differences to copy-structure-related factors, particularly when paired with expression- and product-level measurements. Across these analyses, the emphasis is on reproducibility of molecular population composition under comparable conditions, rather than on a single measurement endpoint.
The molecular basis of high-performance CHO stable cell lines can be summarized in three determinant categories. First, genomic integration architecture and position effects shape transcriptional accessibility and contribute to clone-to-clone variability and expression drift. Second, copy number amplification and expression burden under GS/DHFR selection systems influence expression flux while imposing constraints linked to host processing capacity. Third, epigenetic states govern long-term expression persistence and contribute to silencing risk during serial passaging. Establishing consistent genetic, expression, and protein characterization metrics is essential for mechanistic interpretation of both “high performance” and “stability,” and for conducting meaningful comparisons among clones under controlled analytical conditions.